Goals: create a unified interface for regression standardization to obtain estimates of causal effects such as the average treatment effect, or relative treatment effect.
stdReg2 and stdRegstdReg2 is the next generation of stdReg.
If you are happy using stdReg, you can continue using it
and nothing will change in the near future. With stdReg2 we
aim to solve similar problems but with nicer output, more available
methods, the possibility to include new methods, and mainly to make
maintenance and updating easier.
stdReg2 is available on CRAN and can be installed
with:
install.packages("stdReg2")You can install the development version of stdReg2 from
GitHub with:
# install.packages("remotes")
# remotes::install_github("sachsmc/stdReg2")This is a basic example which shows you how to use regression standardization in a logistic regression model to obtain estimates of the causal risk difference and causal risk ratio:
library(stdReg2)
# basic example
# need to correctly specify the outcome model and no unmeasured confounders
# (+ standard causal assumptions)
set.seed(6)
n <- 100
Z <- rnorm(n)
X <- rnorm(n, mean = Z)
Y <- rbinom(n, 1, prob = (1 + exp(X + Z))^(-1))
dd <- data.frame(Z, X, Y)
x <- standardize_glm(
formula = Y ~ X * Z,
family = "binomial",
data = dd,
values = list(X = 0:1),
contrasts = c("difference", "ratio"),
reference = 0
)
x
#> Outcome formula: Y ~ X * Z
#> <environment: 0x64d9875a9fd0>
#> Outcome family: quasibinomial
#> Outcome link function: logit
#> Exposure: X
#>
#> Tables:
#> X Estimate Std.Error lower.0.95 upper.0.95
#> 1 0 0.519 0.0633 0.395 0.643
#> 2 1 0.391 0.0916 0.211 0.570
#>
#> Reference level: X = 0
#> Contrast: difference
#> X Estimate Std.Error lower.0.95 upper.0.95
#> 1 0 0.000 0.0000 0.000 0.00000
#> 2 1 -0.129 0.0667 -0.259 0.00222
#>
#> Reference level: X = 0
#> Contrast: ratio
#> X Estimate Std.Error lower.0.95 upper.0.95
#> 1 0 1.000 0.000 1.000 1.00
#> 2 1 0.752 0.132 0.494 1.01
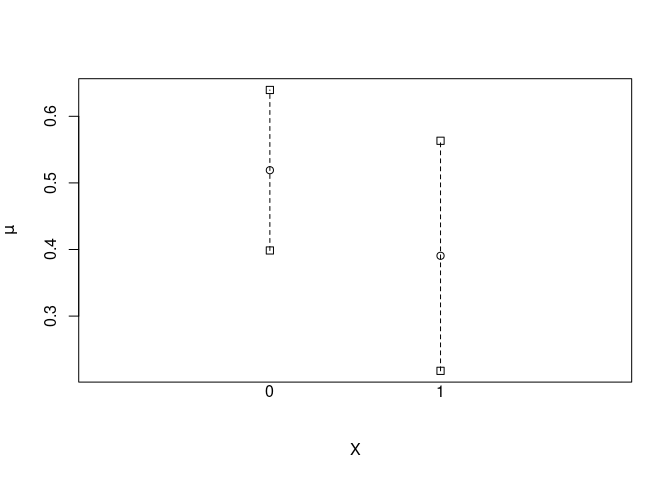
plot(x)
tidy(x)
#> X Estimate Std.Error lower.0.95 upper.0.95 contrast transform
#> 1 0 0.5190639 0.06327946 0.3950384 0.643089345 none identity
#> 2 1 0.3905311 0.09158654 0.2110248 0.570037429 none identity
#> 3 0 0.0000000 0.00000000 0.0000000 0.000000000 difference identity
#> 4 1 -0.1285328 0.06671133 -0.2592846 0.002219041 difference identity
#> 5 0 1.0000000 0.00000000 1.0000000 1.000000000 ratio identity
#> 6 1 0.7523758 0.13178815 0.4940758 1.010675840 ratio identityFor more detailed examples, see the vignette “Estimation of causal effects using stdReg2”.
citation("stdReg2")
#> To cite package 'stdReg2' in publications use:
#>
#> Sachs, C M, Ohlendorff, Sebastian J, Brand, Adam, Sj"olander, Arvid,
#> Gabriel, E E (2025). "The next generation of regression
#> standardization with the R package stdReg2." _Annals of
#> Epidemiology_, *105*, 66-74. doi:10.1016/j.annepidem.2025.04.002
#> <https://doi.org/10.1016/j.annepidem.2025.04.002>.
#>
#> A BibTeX entry for LaTeX users is
#>
#> @Article{,
#> title = {The next generation of regression standardization with the R package stdReg2},
#> author = {{Sachs} and Michael C and {Ohlendorff} and Johan Sebastian and {Brand} and {Adam} and {Sj{"o}lander} and {Arvid} and {Gabriel} and Erin E},
#> journal = {Annals of Epidemiology},
#> volume = {105},
#> pages = {66--74},
#> year = {2025},
#> publisher = {Elsevier},
#> doi = {https://doi.org/10.1016/j.annepidem.2025.04.002},
#> }